MOOZY is a foundation model for computational pathology that treats the patient case, not the individual slide, as the fundamental unit of representation. It encodes one or more whole-slide images (WSIs) into a single 768-dimensional case-level embedding that captures dependencies across all slides from the same patient. Trained entirely on public data with 85.8M parameters (14x smaller than GigaPath), MOOZY outperforms larger models on classification tasks across diverse organs and cancer types.
- News
- Quick Start
- Method Overview
- Training
- Results
- Notes from the Authors
- Acknowledgment
- Citation
- Contact
- License
- [2026/04] MOOZY is now public! Check out the announcements on X (Twitter) and LinkedIn.
pip install moozyModel weights download automatically on first use. No access gates, no manual downloads, no HuggingFace approval.
# Encode a patient case from pre-extracted H5 feature files
moozy encode slide_1.h5 slide_2.h5 --output case_embedding.h5
# Encode directly from raw whole-slide images
moozy encode slide_1.svs slide_2.svs --output case_embedding.h5Or use the Python API:
from moozy.encoding import run_encoding
run_encoding(
slide_paths=["slide_1.h5", "slide_2.h5"],
output_path="case_embedding.h5",
)The output H5 file contains a 768-d case-level embedding ready for downstream tasks: classification, survival prediction, or retrieval.
All encoding arguments (data, runtime, raw WSI options, mixed precision) are documented in docs/encode.md.
conda create -n moozy python=3.12 -y
conda activate moozy
pip install moozyvenv
python -m venv moozy-env
source moozy-env/bin/activate
pip install moozyuv
uv venv moozy-env
source moozy-env/bin/activate
uv pip install moozyThe output is a standard H5 file. Load it with h5py:
import h5py
with h5py.File("case_embedding.h5", "r") as f:
embedding = f["features"][:] # (768,) float32 case-level embedding
# Use the embedding for downstream tasks
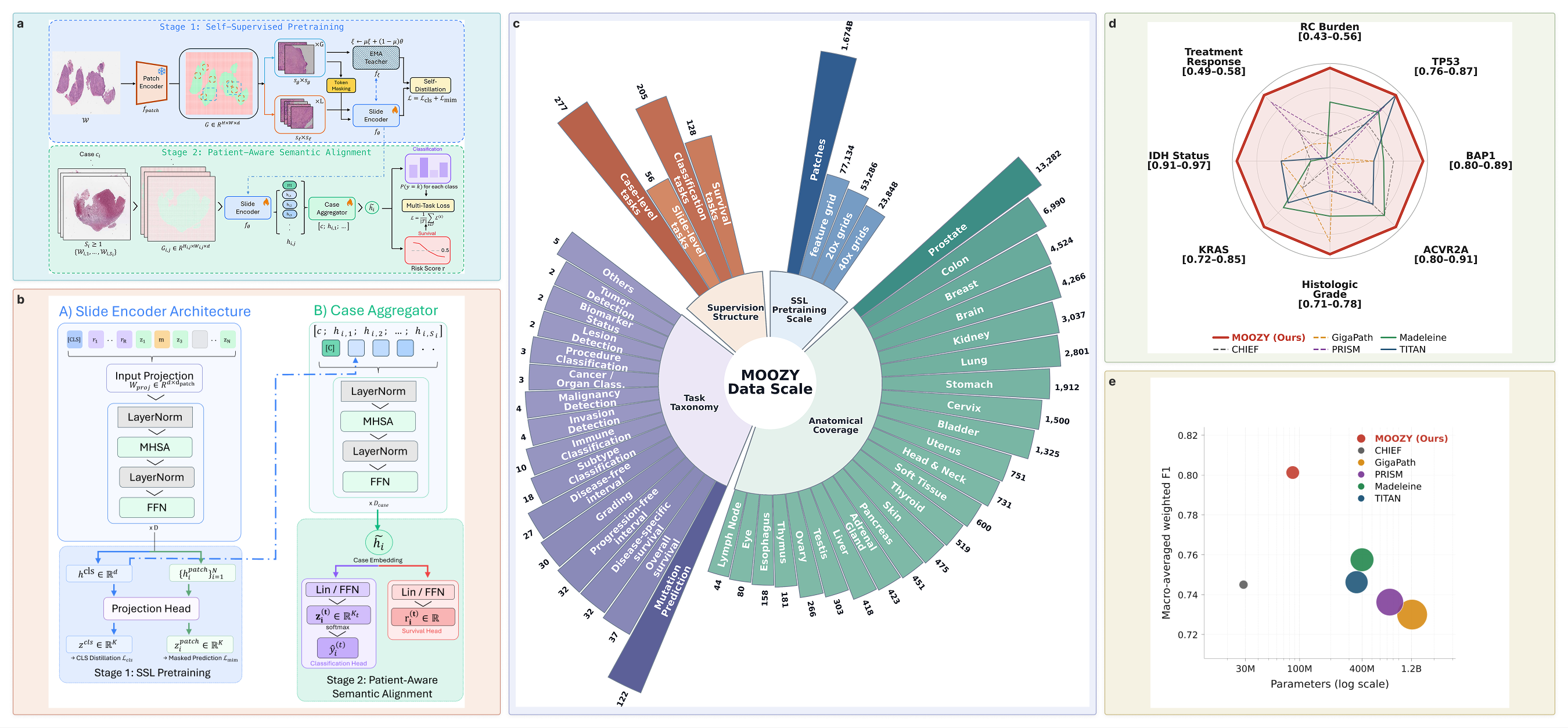
# e.g., as input to a linear probe, k-NN, MLP probe, or clusteringMOOZY is a two-stage pipeline that first learns slide-level representations through self-supervised learning, then aligns them with clinical meaning through multi-task supervision.
Stage 1: Self-supervised slide encoder. A vision transformer learns context-aware spatial representations from 77,134 unlabeled public histopathology slides (~1.67 billion patches across 23 anatomical sites) using masked self-distillation. No labels are used. The slide encoder captures tissue morphology, spatial context, and inter-region relationships across the whole slide.
Stage 2: Patient-aware multi-task alignment. The pretrained slide encoder is fine-tuned end-to-end with a case transformer that models dependencies across all slides from the same patient. A learnable [CASE] token aggregates per-slide embeddings into a single case-level representation. Multi-task supervision across 333 tasks (205 classification, 128 survival) from 56 public datasets provides broad clinical grounding. All task heads are discarded after training, leaving a general-purpose patient encoder.
For detailed model specifications, see the model card.
Both training stages are fully open-source and reproducible using only public data. All training arguments (data, model, optimization, checkpointing, logging, runtime) are documented in the Stage 1 and Stage 2 training docs.
For local multi-GPU training, use the launch scripts in scripts/:
# Stage 1: Self-supervised pretraining
GPU_IDS=0,1,2,3,4,5,6,7 bash scripts/train_stage1.sh
# Stage 2: Multi-task alignment
GPU_IDS=0,1,2,3,4,5,6,7 bash scripts/train_stage2.shSLURM job templates are provided in slurm/ for cluster environments:
| Script | Description |
|---|---|
slurm/single_gpu.sh |
Single-GPU training |
slurm/multi_gpu.sh |
Multi-GPU training on one node |
slurm/multi_node.sh |
Multi-node distributed training |
slurm/inference.sh |
Patient encoding |
Frozen-feature MLP probe comparison against slide encoder baselines on eight held-out tasks. Bold indicates the best result per metric.
| Task | Metric | CHIEF | GigaPath | PRISM | Madeleine | TITAN | MOOZY |
|---|---|---|---|---|---|---|---|
| Residual Cancer Burden | F1 | 0.46 | 0.45 | 0.46 | 0.51 | 0.43 | 0.56 |
| AUC | 0.60 | 0.55 | 0.58 | 0.63 | 0.58 | 0.74 | |
| Bal Acc | 0.44 | 0.40 | 0.43 | 0.48 | 0.38 | 0.51 | |
| TP53 Mutation | F1 | 0.82 | 0.76 | 0.85 | 0.84 | 0.87 | 0.87 |
| AUC | 0.81 | 0.76 | 0.85 | 0.85 | 0.91 | 0.86 | |
| Bal Acc | 0.83 | 0.76 | 0.84 | 0.84 | 0.88 | 0.86 | |
| BAP1 Mutation | F1 | 0.86 | 0.84 | 0.80 | 0.85 | 0.84 | 0.89 |
| AUC | 0.75 | 0.63 | 0.71 | 0.78 | 0.82 | 0.79 | |
| Bal Acc | 0.75 | 0.66 | 0.66 | 0.75 | 0.75 | 0.78 | |
| ACVR2A Mutation | F1 | 0.89 | 0.80 | 0.85 | 0.89 | 0.87 | 0.91 |
| AUC | 0.80 | 0.74 | 0.83 | 0.76 | 0.79 | 0.91 | |
| Bal Acc | 0.80 | 0.65 | 0.81 | 0.81 | 0.76 | 0.90 | |
| Histologic Grade | F1 | 0.71 | 0.77 | 0.73 | 0.75 | 0.73 | 0.78 |
| AUC | 0.71 | 0.77 | 0.67 | 0.74 | 0.71 | 0.75 | |
| Bal Acc | 0.73 | 0.77 | 0.73 | 0.74 | 0.73 | 0.77 | |
| KRAS Mutation | F1 | 0.77 | 0.77 | 0.72 | 0.81 | 0.80 | 0.85 |
| AUC | 0.76 | 0.72 | 0.61 | 0.70 | 0.80 | 0.80 | |
| Bal Acc | 0.74 | 0.76 | 0.63 | 0.77 | 0.81 | 0.79 | |
| IDH Status | F1 | 0.92 | 0.94 | 0.91 | 0.92 | 0.94 | 0.97 |
| AUC | 0.96 | 0.97 | 0.95 | 0.96 | 0.97 | 0.99 | |
| Bal Acc | 0.92 | 0.94 | 0.91 | 0.91 | 0.94 | 0.97 | |
| Treatment Response | F1 | 0.53 | 0.51 | 0.57 | 0.49 | 0.49 | 0.58 |
| AUC | 0.70 | 0.68 | 0.69 | 0.59 | 0.60 | 0.68 | |
| Bal Acc | 0.48 | 0.40 | 0.51 | 0.35 | 0.37 | 0.48 |
Mean values from five-fold frozen-feature evaluation. Full results are in the paper.
Across all eight tasks, MOOZY improves macro averages over TITAN by +7.4% weighted F1, +5.5% AUC, and +7.8% balanced accuracy, and over PRISM by +8.8% F1, +10.7% AUC, and +9.8% balanced accuracy, with 14x fewer parameters than GigaPath.
A few readers have asked us why the main tables in the paper use a non-linear (MLP) probe rather than the more conventional linear probe. We wanted to share the reasoning here.
Slide-encoder embeddings are not guaranteed to be linearly separable. Some encoders (e.g. contrastive or aligned multimodal models) are explicitly trained to structure features along linear axes, while others organize information through higher-order interactions that a linear classifier cannot access. A linear probe rewards the former and can underrepresent the latter even when both carry the same useful information. We chose the MLP probe as the primary benchmark because it treats every encoder symmetrically. The classifier is free to use whichever structure is present in the features, without requiring it to be linearly separable. In pathology, clinically relevant phenotypes depend on nonlinear mixtures of cellular morphology and its spatial context, so a linear head on top of frozen slide features is expected to leave real signal unread. Linear-probe results still matter, since they are a more conservative measure of how features transfer to downstream pipelines that use a simple logistic-regression head, so we report both.
The full linear-probe version of the slide-encoder comparison (L2-regularized multinomial logistic regression on the same frozen features) is reported in the appendix of our paper, alongside the per-task breakdowns. MOOZY's numbers drop when the classifier is swapped from an MLP head to a linear head, but that is true of every slide encoder we evaluated, not just MOOZY. Averaged across all six encoders (CHIEF, GigaPath, PRISM, Madeleine, TITAN, and MOOZY), the mean macro-average loss when moving from MLP to linear is about 0.097 on weighted F1, 0.027 on weighted ROC-AUC, and 0.087 on balanced accuracy. The much smaller drop on ROC-AUC is consistent with the rest of this note. A linear head preserves the ordering of predictions across the board but loses the clean decision boundary that an MLP head can find.
The same linear-vs-MLP question can also be asked against the patch-encoder plus trained-MIL baselines. In the table below, each non-MOOZY row pairs a frozen patch encoder with a task-specific MIL aggregator trained from scratch (MeanMIL, ABMIL, CLAM, DSMIL, TransMIL) and averages across the five architectures. The Backbone row uses the same ViT-S/8 Lunit DINOv2 patch encoder that MOOZY itself uses internally (Kang et al. 2023), so this row isolates what MOOZY's slide and case encoder add on top of the shared patch features.
Linear classifier on MOOZY vs. trained MIL aggregators (macro average over 5 MIL architectures).
| Patch encoder | Weighted F1 | Weighted ROC-AUC | Balanced Accuracy |
|---|---|---|---|
| Backbone (MOOZY's patch encoder) | 0.733 | 0.735 | 0.686 |
| UNI v2 | 0.716 | 0.719 | 0.660 |
| Phikon v2 | 0.715 | 0.724 | 0.654 |
| CONCH v1.5 | 0.746 | 0.751 | 0.696 |
| MUSK | 0.729 | 0.725 | 0.679 |
| MOOZY (linear probe) | 0.698 | 0.778 | 0.674 |
Macro averages across the eight held-out tasks. Non-MOOZY rows use frozen patch features with a trained MIL aggregator head, averaged over five architectures. MOOZY uses a linear classifier on top of its frozen case embedding, with no MIL training.
MOOZY's slide and case encoder add real signal on top of the shared patch features, but that signal is non-linearly structured. A linear head recovers the ordering but not the decision boundary, so MOOZY keeps the top ROC-AUC under the linear probe (+0.027 over CONCH v1.5) while trailing CONCH v1.5 on weighted F1 and balanced accuracy. The Backbone row tells the same story from a different angle, since its frozen features are the same ones MOOZY is built on. Under MLP, the Backbone-to-MOOZY gap is +0.068 F1, +0.080 ROC-AUC, and +0.072 balanced accuracy. Under linear, only the +0.043 ROC-AUC lift survives, and F1 and balanced accuracy fall by -0.035 and -0.012.
This work was supported by NSERC-DG RGPIN-2022-05378 [M.S.H], Amazon Research Award [M.S.H], and Gina Cody RIF [M.S.H], FRQNT scholarship [Y.K]. Computational resources were provided in part by Calcul Québec and the Digital Research Alliance of Canada.
If you find MOOZY useful, please cite:
@misc{kotp2026moozypatientfirstfoundationmodel,
title={MOOZY: A Patient-First Foundation Model for Computational Pathology},
author={Yousef Kotp and Vincent Quoc-Huy Trinh and Christopher Pal and Mahdi S. Hosseini},
year={2026},
eprint={2603.27048},
archivePrefix={arXiv},
primaryClass={cs.CV},
url={https://arxiv.org/abs/2603.27048},
}For questions, bug reports, or just to say hi, my inbox is open at yousefkotp@outlook.com. I am a human who reads every email, even the ones that start with "I know you're probably busy, but...". Feel free to reach out about anything related to MOOZY, computational pathology, or just to chat about deep learning and its applications in medicine. I also welcome feedback on the codebase and any suggestions for improvement.
This project is licensed under CC BY-NC-SA 4.0.